Table of Contents
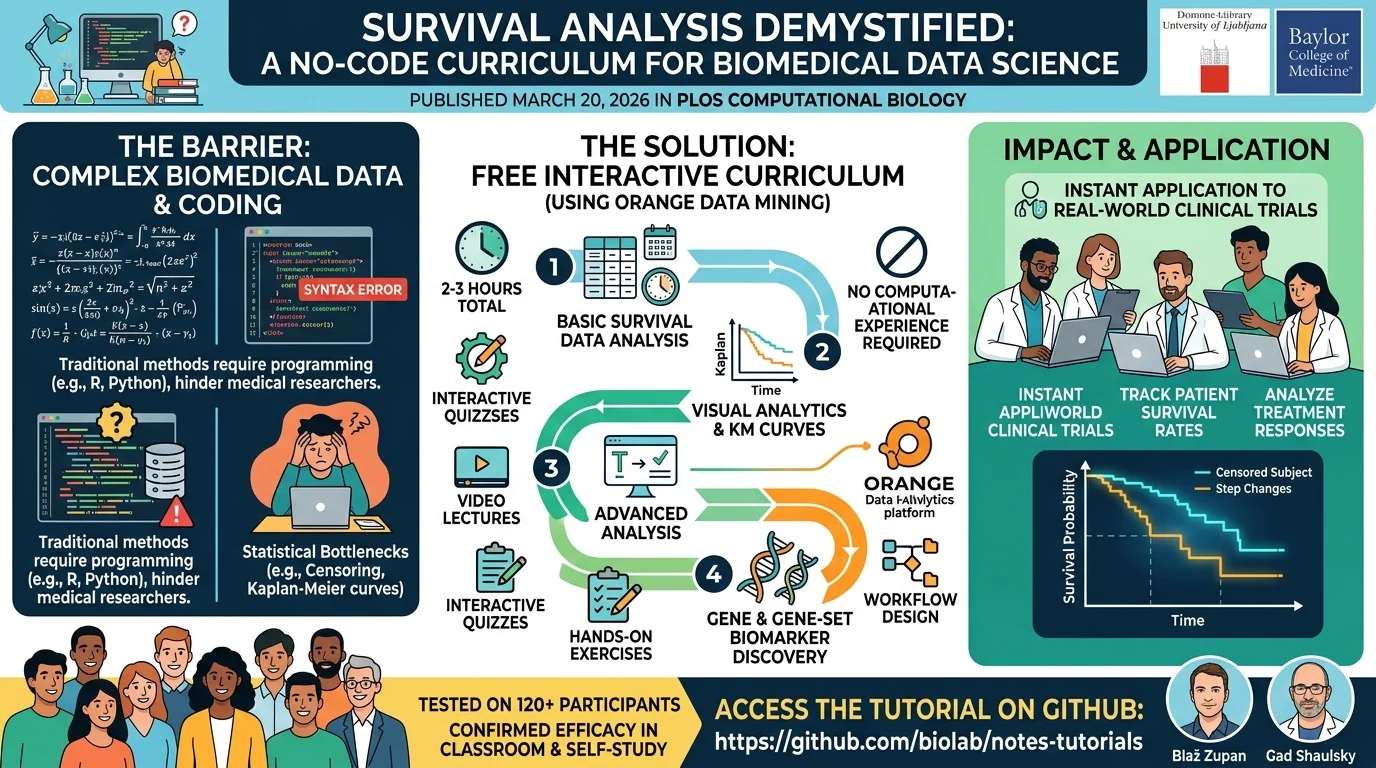
A newly published survival analysis tutorial is breaking down the complex mathematics of biomedical data science into an accessible, no-code format. Released on March 20, 2026, in PLOS Computational Biology, a joint research team from the University of Ljubljana and Baylor College of Medicine introduced a free, interactive curriculum designed specifically for biomarker discovery. This initiative aims to eliminate the programming barriers that often hinder medical students and researchers from mastering time-to-event data analysis.
Designed primarily for biomedical students, educators, and self-learners with zero prior computational experience, this curriculum transforms how clinical data is interpreted. By utilizing visual analytics instead of raw coding, learners can immediately apply these techniques to real-world clinical trials. This enables them to track patient survival rates and treatment responses without getting bogged down by syntax errors or complex software environments.
Survival analysis is a cornerstone of modern medical research, used in thousands of biomedical studies annually to measure the time until a specific event occurs, such as disease progression or patient mortality. However, traditional teaching methods struggle to explain non-intuitive concepts like "censored" subjects - patients who drop out or do not experience the event during the study. Misinterpreting these nuances, or misreading the distinct steps of Kaplan-Meier curves, can lead to flawed clinical conclusions and compromised research outcomes.
To solve this educational bottleneck, the researchers built their curriculum around Orange Data Mining, an open-source, no-code visual analytics platform. The tutorial spans four pedagogical units, requiring only two to three hours to complete. It progresses logically from basic survival data analysis to advanced gene and gene-set biomarker discovery, ensuring a gentle learning curve for all participants.
The comprehensive package goes beyond a standard online guide. It integrates video lectures, written literature, interactive quizzes, and practical hands-on exercises. Students are guided through data creation, workflow design, and the biological interpretation of their findings. The tutorial materials are hosted on GitHub at https://github.com/biolab/notes-tutorials.
The research team, including Blaž Zupan and Gad Shaulsky, successfully tested the curriculum with over 120 participants. The results proved its efficacy in both individual study and classroom environments, confirming that complex statistical methods can be taught effectively without relying on traditional programming languages.
My Take
The introduction of this no-code survival analysis tutorial represents a crucial shift in biomedical education. By leveraging the Orange Data Mining platform, the researchers have effectively decoupled statistical literacy from programming proficiency. The fact that this curriculum was successfully validated with over 120 participants suggests a high demand for visual-first learning tools in medical academia.
As data science becomes increasingly central to clinical research, open-access, intuitive frameworks like this will be essential. They not only accelerate biomarker discovery but also reduce analytical bottlenecks in laboratories, allowing researchers to focus on biological insights rather than software troubleshooting.
Frequently Asked Questions
Who is the target audience for this survival analysis tutorial?
It is designed for biomedical students, researchers, and educators, particularly those without prior programming or advanced computational experience.
What software is required to complete the exercises?
The curriculum utilizes Orange Data Mining, a free, open-source visual analytics platform that requires no coding.
How long does it take to complete the course?
The tutorial is structured into four units and is designed to be completed in approximately two to three hours.